Note: this is a post to hold the R script that is referred to in this post. It’s even more awfully technical than the post that refers to it; readers not interested or familiar with R code or this type of analysis may want to skip it!
You’d probably want to copy this all in to an R script (and perhaps view in r studio) to make sense of it.
# Disturbance by patch to determine landscape predictors of disturbance —-
# Housekeeping —-
setwd(“C:/Users/ke/Dropbox/PhD/Chapter 3 – spatial analysis/R”)
library(lme4); library(ggplot2); library(plyr); library(lattice); library(car); require(lmerTest); library(nlme)
source(“HighstatLibV6.R”)
dp
# Start of data exploration —-
names(dp)
str(dp)
summary(dp)
attach(dp)
head(dp)
sum(is.na(dp)) # no missing values
# 4.2 looking for outliers
vars1 “greenstone” , “esa” , “cons_estate” , “iron_formation”)
Mydotplot(dp[, vars1])
vars2 “mining_activity”,”commodity”,”wheatbelt_dist”,”town_dist”,”patch_area”,”disturbed.area”, “dist_perc”)
Mydotplot(dp[, vars2])
vars3 Mydotplot(dp[, vars3])
# no outliers once the tenement years issue was fixed (in raw data), but some vars need transformation
# Transforming variables —-
# mini-tutorial on how scaling works:
x s.x s.x # the resultant scaled variable stored the scaling attributes: the centre (mean) and scale (multiplier)
attr(s.x,”scaled:center”) # 5.5; extracting the centre, the mean of the original vector, from which each value was subtracted to do the scaling
attr(s.x,”scaled:scale”) # 3.02765; extracting the scale (the multiplier)
x.recreated = s.x*attr(s.x,’scaled:scale’) + attr(s.x,’scaled:center’) # to unscale just multiply the scaled vector by the ‘scale’ and add the ‘centre’
# transforming disturbance percentage:
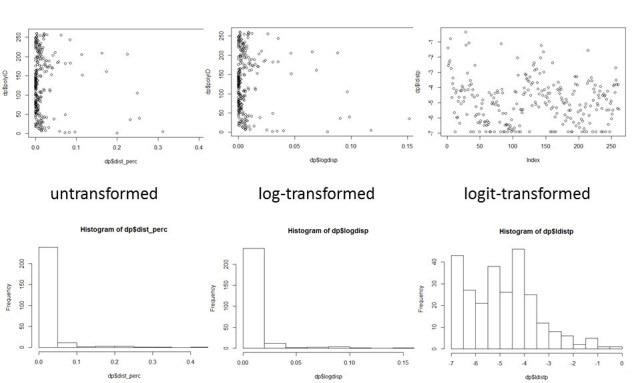
plot(dp$polyID~dp$dist_perc)
hist(dp$dist_perc) # too skewed
dist_perc1 = dist_perc + 1
dp$logdisp = log(dist_perc1)
range(dp$logdisp)
hist(dp$logdisp)
plot(dp$polyID~dp$logdisp) # still too squashed. try log10 transformation?
dp$logdisp = log10(dist_perc1)
attach(dp)
range(dp$logdisp)
hist(dp$logdisp)
plot(dp$polyID~dp$logdisp) # still too skewed. Let’s try a logit transformation:
qlogis(0.9) # Getting familiar w logit transformations… this returns 2.197. qlogis does a logit transformation on (a set of) values.
plogis(2.197225) # returns 0.9. plogis ‘undoes’ the logit transformation.
dp$ldistp = (qlogis(dp$dist_perc+0.001)) # ldistp is logit transformation of the percent disturbed plus 0.001
plot(dp$ldistp) # Looks good.
range(dp$ldistp)
hist(dp$ldistp)
dp$s.dist = scale(dp$ldistp) # s.dist is the logit-transformed, scaled disturbed percentage
hist(dp$s.dist)
# OK I need to either do a logistic regression here, or simply logit-transform the response before modelling linearly
# doing a logistic transformation and then modelling linearly – advised by Michael Renton
# transforming and scaling ten_yrs:
range(ten_yrs) # the 177 is a valid value.
plot(ten_yrs) # needs some transformation
dp$lntyrs = log((ten_yrs+1))
plot(dp$lntyrs) # yep the natural log looks good.
range(dp$lntyrs)
dp$s.lntyrs = scale(dp$lntyrs, center = TRUE, scale = TRUE)
plot(dp$lntyrs)
plot(dp$s.lntyrs)
# transforming patch area:
range(dp$patch_area)
dp$patchkm2 = dp$patch_area/1000000 # make kms instead of m2
range(dp$patchkm2)
hist(dp$patchkm2)
dp$ln.pkm = log(dp$patchkm2) # natural log of patch area
hist(dp$ln.pkm) # centred and normalish
# transforming minedex_count:
plot(minedex_count)
table(dp$minedex_count)
plot(log(minedex_count+1)) # somewhat better
logmx = log(minedex_count+1)
range(logmx)
dp$s.mx = scale(logmx)
plot(dp$s.mx)
# transforming distance to wheatbelt:
range(wheatbelt_dist)
dp$wbkm = wheatbelt_dist/1000
plot(dp$wbkm)
range(dp$wbkm)
dp$s.wb = scale(dp$wbkm)
plot(dp$s.wb)
# transforming distance to town:
dp$tnkm = town_dist/1000
plot(dp$tnkm)
range(dp$tnkm)
dp$s.tn = scale(dp$tnkm)
plot(dp$s.tn)
# not much can be done with the commodity variable:
summary(dp$commodity) # not sure if having the “no commodity stated” value is useful…
# dp$commodity[dp$commodity==”no commodity stated”] <- “none” # creates NAs
# adding second-order polynomials of the numerical variables
dp$tenyrs2 = dp$s.lntyrs^2
dp$mx2 = dp$s.mx^2
dp$wb2 = dp$s.wb^2
dp$tn2 = dp$s.tn^2
range(dp$s.lntyrs)
range(dp$tenyrs2)
# combining greenstone and ironstone lithology data into one categorical variable
dp$lithology = ifelse(greenstone == “greenstone”,”greenstone”,”other.lithology”)
dp$iron_formation
dp$lithology = ifelse(iron_formation == “iron-form”,”iron.formation”,dp$lithology)
dp$lithology
table(dp$lithology) # clever duck
# Looking for collinearity —-
pairs(dp[,vars1], lower.panel = panel.cor)
#i.e. SA number, SA_mines and category all related – use SA_mines
pairs(dp[,vars2], lower.panel = panel.cor)
miningvars = c(“SA_mines”,”greenstone”,”iron_formation”,”minedex_count”,”ten_yrs”,”mining_activity”,”commodity”)
pairs(dp[,miningvars], lower.panel = panel.cor)
# also strong collinearity between minedex count and dt-count, mined and
# note mining activity replaces expl_ten and mined.
# low 0.4 collinearity between minedex count and ten_yrs, and 0.5 between SA_mines and mining activity
summary(dp$dist_perc ~ dp$veg_form)
summary(dp$cons_estate)
dpvars = c(“pastoral”,”greenstone”,”esa”,”cons_estate”,”iron_formation”,”sched_1″,”veg_form”,
“mining_activity”,”commodity”,”s.mx”,”s.lntyrs”,”s.wb”,”s.tn”,”s.dist”)
Mydotplot(dp[, dpvars])
pairs(dp[,dpvars], lower.panel = panel.cor)
# ok, have dealt with collinearity. There’s nothing above 0.5, and even the 0.5 is not important
# looking at possible X Y relationships —-
# first the numeric variables: plot Y against the Xes and calculate pearson coefficients:
dpnumvars = c(“s.mx”,”s.lntyrs”,”s.wb”,”s.tn”,”s.dist”)
pairs(dp[,dpnumvars], lower.panel = panel.cor)
# there appears to be a positive relationship between the area disturbed and both the
# number of mining projects and the total duration of tenements in an area.
# No relation with distance from wheatbelt, and slight negative relationship with distance from town,
# which makes sense – areas near towns are more likely to get more disturbed and to stop tracks from growing over etc.
plot(dp$dist_perc~dp$lntyrs) #remember that dp$lntyrs = log((ten_yrs+1))
plot(dp$ldistp~dp$lntyrs)
#then categorical variables:
boxplot(dp$s.dist~dp$pastoral) # no real effect of pastoral activity
boxplot(dp$s.dist~dp$greenstone) # more disturbance on greenstone
boxplot(dp$s.dist~dp$esa) # more disturbance on ESAs
boxplot(dp$s.dist~dp$cons_estate) # greatest disturbance on former leasehold then class As!
# UCL has same mean distrubance as gazetted conservation estate; though higher variance.
boxplot(dp$s.dist~dp$iron_formation) # more disturbance on BIFs
boxplot(dp$s.dist~dp$sched_1) # more disturbance on schedule 1 areas
boxplot(dp$s.dist~dp$veg_form) # less disturbance in bare areas and succulent steppe
boxplot(dp$dist_perc~dp$veg_form)
boxplot(dp$s.dist~dp$mining_activity) # more disturbance where there’s been mining tenement.
# strangely similar between exploration and no mining tenement.
boxplot(dp$s.dist~dp$commodity) # most disturbance around gold mining projects, maybe not significant
# Looking for interactions —-
coplot(s.dist ~ s.mx | factor(commodity), data = dp)
# R displays from lower left, across, to upper right.
coplot(s.dist ~ s.mx | factor(commodity), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
# there could be an interaction between the commodity and the number of mining projects
coplot(s.dist ~ s.lntyrs | factor(pastoral), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
# There could be an interaction between tenement years and pastoral, same as found in regional-scale analysis.
coplot(s.dist ~ factor(greenstone) | factor(pastoral), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
# maybe nothing between greenstone and pastoral
coplot(s.dist ~ factor(veg_form) | factor(greenstone), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
# maybe nothing between greenstone and pastoral
coplot(s.dist ~ s.tn | factor(veg_form), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
# well there doesn’t seem to be much mallee near towns; looks like there could be an interaction
coplot(s.dist ~ s.tn | factor(esa), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
# definitely looks like an interaction between esas and dist to town
coplot(s.dist ~ s.wb | factor(esa), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
# same again – interaction between esas and distance to wheatbelt, although esas tend to be much closer to wheatbelt than non-esas.
coplot(s.dist ~ factor(veg_form) | factor(esa), data = dp,
panel = function(x, y, …) {
tmp abline(tmp)
points(x, y) })
coplot(s.dist ~ factor(mining_activity) | factor(veg_form), data = dp )
# most of the mining and exploration appears to be in woodland
# Fitting and selecting models —-
# 4.7.1 using gaussian family and logit link with lme4
# glmer example: model1 dp1 = glmer(dist_perc ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+
(1|SA_number),
data=dp, family = gaussian (link = ‘logit’))
summary(dp1)
E1 F1 plot(x = F1,
y = E1,
xlab = “Fitted values”,
ylab = “Residuals”,
main = “Homogeneity?”)
abline(h = 0, v = 0, lty = 2)
plot(dp1)
plot(F1)
# 4.7.2 Fitting a linear mixed model with a logit-transformed response in lmerTest
dp2 = lmer(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+mx2+wb2+tn2+
(1|SA_number), REML=TRUE,
data=dp)
summary(dp2)
plot(dp2)
summary(dp$commodity)
# dp$commodity[is.na(dp$commodity)] <- “none”
# compare with model that doesn’t have random effects to see if they are important
dp2.gls = gls(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+mx2+wb2+tn2,
data=dp)
summary(dp2.gls)
plot(dp2.gls)
anova(dp2.gls,dp2)
anova(dp2)
plot(dp2)
E2 F2 plot(x = F2,
y = E2,
xlab = “Fitted values”,
ylab = “Residuals”,
main = “Homogeneity?”)
abline(h = 0, v = 0, lty = 2)
table(dp$SA_number)
length(E2)
dp3 = lmer(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+mx2+wb2+tn2+
(1|SA_number),
data=dp, REML = FALSE)
summary(dp3)
anova(dp3)
dp4 = lmer(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+
(1|SA_number),
data=dp, REML = FALSE)
summary(dp4)
anova(dp4)
dp5 = lmer(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(veg_form)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+
(1|SA_number),
data=dp, REML = FALSE)
summary(dp5)
anova(dp5)
dp6 = lmer(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(veg_form)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+
(1|SA_number),
data=dp, REML = FALSE)
summary(dp6)
anova(dp6)
# 4.7.3 Fitting a linear mixed model with a logit-transformed response in nlme
library(nlme)
# first check the significance of the random effect structure:
dp1 = lme(s.dist ~ factor(pastoral)+factor(iron_formation)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(sched_1)+factor(veg_form) +factor(commodity)+factor(mining_activity)+
s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+mx2+wb2+tn2+ln.pkm,
random = ~1|SA_number,method = “REML”,
data=dp)
summary(dp1)
plot(dp1)
# compare with model that doesn’t have random effects to see if they are important
dp1.gls = gls(s.dist ~ factor(pastoral) +factor(iron_formation) +factor(greenstone) +factor(esa) +
factor(cons_estate) +factor(sched_1) +factor(veg_form) +factor(commodity) +
factor(mining_activity) +s.lntyrs +s.mx +s.wb +s.tn +tenyrs2 +mx2 +wb2 +tn2 +ln.pkm,
data=dp)
dp$s.dist
summary(dp1.gls)
plot(dp1.gls)
anova(dp1.gls,dpn1)
# The p-value of the Maximum Likelihood ratio test comparison between the mixed and fixed methods
# (both calculated using REML, which allows us to apply the likelihood ratio test) is 0.0154,
# indicating that the random factor is significant.
# check the model:
E2 F2 op MyYlab <- “Residuals”
plot(x = F2, y = E2, xlab = “Fitted values”, ylab = MyYlab, main=”Residuals versus fitted”)
boxplot(E2 ~ pastoral, data = dp,
main = “Pastoral tenure”, ylab = MyYlab)
boxplot(E2 ~ greenstone, data = dp,
main = “Greenstone lithology”, ylab = MyYlab)
plot(x = dp$s.lntyrs, y = E2, ylab = MyYlab,
main = “Tenement duration”, xlab = “scaled log of total tenement duration”)
par(op)
#haven’t checked against all variables but there’s too many, moving on for now and will check when model is smaller.
# Next, I’m going to fit the ‘beyond optimal’ model with ML, then reduce it manually
# then validate it,
dp2 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+mx2+wb2+tn2+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp2)
# remove tn2
dp3 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+mx2+wb2+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
anova(dp2,dp3)
# remove mx2
dp4 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+wb2+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
anova(dp4,dp3) #yes, wasn’t significant
summary(dp4)
# remove wb2
dp5 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+tenyrs2+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp5)
#remove tenyrs2
dp6 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.mx+s.wb+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp6)
# remove s.mx
dp7 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(sched_1)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.wb+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp7)
# remove schedule 1 areas
dp8 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(iron_formation)+factor(veg_form)+factor(commodity)+
factor(mining_activity)+s.lntyrs+s.wb+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp8)
# remove iron_form
dp9 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+factor(mining_activity)+s.lntyrs+s.wb+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp9)
# remove either distance to wheatbelt or mining activity
dp9.no.wb = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+factor(mining_activity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
dp9.no.ma = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+s.lntyrs+s.wb+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
anova(dp9,dp9.no.wb) # p = 0.1054
anova(dp9,dp9.no.ma) # p = 0.2048
# so remove mining activity
dp10 = lme(s.dist ~ factor(pastoral)+factor(greenstone)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+s.lntyrs+s.wb+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp10)
# not significant are: greenstone, cons estate, wheatbelt. try remove each, one at a time.
dp10.no.green dp10.no.cons dp10.no.wb
anova(dp10,dp10.no.green) # p = 0.0539
anova(dp10,dp10.no.cons) # p < 0.0001
anova(dp10,dp10.no.wb) # p = 0.0395
# got to remove greenstone
dp11 = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+s.lntyrs+s.wb+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp11)
# pastoral is significant
# esa is significant
# cons estate need to be checked but is probably highly significant
# veg form looks significant
# commodity looks significant
# tenement years is highly significant
# s.wb – what is that still doing here? the p values from the model are so different from anova comparing models!
dp11.no.wb = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
anova(dp11,dp11.no.wb)
# yep get rid of wheatbelt
dp12 = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp12)
# I think we’re there so just double checking the significance of every variable:
dp12.no.pastoral = lme(s.dist ~ factor(esa)+factor(cons_estate)+factor(veg_form)+factor(commodity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”, data=dp)
dp12.no.esa = lme(s.dist ~ factor(pastoral)+factor(cons_estate)+factor(veg_form)+factor(commodity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”, data=dp)
dp12.no.cons = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(veg_form)+factor(commodity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”, data=dp)
dp12.no.veg = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(commodity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”, data=dp)
dp12.no.comm = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(veg_form)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”, data=dp)
dp12.no.lntyrs = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(veg_form)+factor(commodity)+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”, data=dp)
dp12.no.tn = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(veg_form)+factor(commodity)+s.lntyrs+ln.pkm,
random = ~1|SA_number,method = “ML”, data=dp)
dp12.no.pkm = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(veg_form)+factor(commodity)+s.lntyrs+s.tn,
random = ~1|SA_number,method = “ML”, data=dp)
anova(dp12,dp12.no.pastoral) # p = 0.0046
anova(dp12,dp12.no.esa) # p = 0.0001
anova(dp12,dp12.no.cons) # p = 0.0001
anova(dp12,dp12.no.veg) # p = 0.0206
anova(dp12,dp12.no.comm) # p = 0.0001
anova(dp12,dp12.no.lntyrs) # p = 0.0001
anova(dp12,dp12.no.tn) # p = 0.000001
anova(dp12,dp12.no.pkm) # p = 0.0348
# all significant, and lower AIC values!
# what if i try to reintroduce greenstone?
dp12.plus.green = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(greenstone)+
factor(veg_form)+factor(commodity)+s.lntyrs+s.tn+ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
anova(dp12,dp12.plus.green)
# greenstone is marginally significant with p = 0.0562 and a slightly lower AIC value (but within 2 AIC). Leave out.
#what about interaction between mining and pastoralism?
dp13 = lme(s.dist ~ factor(pastoral)+factor(esa)+factor(cons_estate)+
factor(veg_form)+factor(commodity)+s.lntyrs+s.tn+ln.pkm+s.lntyrs:factor(pastoral),
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp13)
# interaction not significant
# I’ll also just try to use non-scaled versions of the variables for ease of interpretation if possible:
dp14 = lme(ldistp ~ factor(pastoral)+factor(esa)+factor(cons_estate)+
factor(veg_form) +factor(commodity) +lntyrs +tnkm +ln.pkm,
random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp14)
# excellent! unscaled it is…
# what about removing commodity as it’s not helpful as is, and seeing if this
# and allows for reinstatement of greenstone too, and removal of patch area:
dp15 = lme(ldistp ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(veg_form)+
lntyrs+tnkm+factor(greenstone),random = ~1|SA_number,method = “ML”,
data=dp)
summary(dp15)
#yes, lets just recheck the significance of all variables:
dp15.no.pastoral = update(dp15, .~. – factor(pastoral))
dp15.no.esa = update(dp15, .~. – factor(esa))
dp15.no.cons = update(dp15, .~. – factor(cons_estate))
dp15.no.veg = update(dp15, .~. – factor(veg_form))
dp15.no.green = update(dp15, .~. – factor(greenstone))
dp15.no.lntyrs = update(dp15, .~. – lntyrs)
dp15.no.tn = update(dp15, .~. – tnkm)
dp15.w.pkm = lme(ldistp ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(veg_form)+lntyrs+
tnkm+ln.pkm+factor(greenstone)+ln.pkm,random = ~1|SA_number,method = “ML”,data=dp)
anova(dp15,dp15.no.pastoral) # p = 0.0031
anova(dp15,dp15.no.esa) # p = 0.0004
anova(dp15,dp15.no.cons) # p < 0.0001
anova(dp15,dp15.no.veg) # p = 0.005
anova(dp15,dp15.no.green) # p = 0.006
anova(dp15,dp15.no.lntyrs) # p = 0.0001
anova(dp15,dp15.no.tn) # p = 0.0002
anova(dp15,dp15.w.pkm) # p = 0.778 # excluding patch area is justified.
# otherwise all significant, and lower AIC values!
# Final dp model, using REML and with unscaled response variable
finaldp = lme(ldistp ~ factor(pastoral)+factor(esa)+factor(cons_estate)+factor(veg_form)+
factor(greenstone)+lntyrs+tnkm,random = ~1|SA_number,method = “REML”, data=dp)
summary(finaldp)
# Validate model
par(mfrow=c(2,2))
plot(finaldp)
E1 F1
par(mfrow = c(1, 1))
plot(x = F1,
y = E1,
xlab = “Fitted values”,
ylab = “Residuals”,
main = “Homogeneity?”)
abline(h = 0, v = 0, lty = 2)
# according to Zuur 2009, looks ok
boxplot(E1 ~ factor(dp$greenstone)) # compare variance across factor levels
boxplot(E1 ~ factor(dp$esa))
boxplot(E1 ~ factor(dp$cons_estate))
boxplot(E1 ~ factor(dp$veg_form))
boxplot(E1 ~ factor(dp$commodity))
boxplot(E1 ~ factor(dp$mining_activity))
tapply(E1, INDEX = dp$pastoral, FUN = var)
tapply(E1, INDEX = dp$cons_estate, FUN = var)
# variance varies only by a factor of ~4; Zuur says that it’s a problem if greater than 10. So not a problem.
# could consider removing the outlier
# test for independence:
plot(x = dp$patch_area,
y = E1,
xlab = “patch area”,
ylab = “Residuals”)
abline(h = 0, lty = 2)
cor.test(E1, dp$patch_area) # not significant
# i.e. no clusters above or below; looks ok.
#Normality
hist(E1, main = “Normality”, breaks=10)
#Or qq-plot
qqnorm(E1)
qqline(E1)
# i.e. not so normal
hist(dp$s.dist ) # very skewed no worries
#But plot residuals also versus covariates NOT in the model!
boxplot(E1 ~ s.wb) ; abline(h = 0, lty = 2) # confirms no effect of distance to wheatbelt
boxplot(E1 ~ s.tn) ; abline(h = 0, lty = 2) # no residual effect of distance to town
# Final model and sketch with predictions —-
finaldp = lme(ldistp ~ factor(pastoral) +factor(esa) +factor(cons_estate) +factor(veg_form) +
factor(greenstone)+lntyrs+tnkm,random = ~1|SA_number,method = “REML”, data=dp)
summary(finaldp)
summary(finaldp)$coef
anova.lme(finaldp)
# I tried many different ways to predict from the model using a newdata dataframe but couldn’t get it to work.
# here i use the values i calculated manually in excel using the model outputs:
# remember how to un-logit the response variable:
plogis(2.197225) # returns 0.9. plogis undoes the logit transformation.
ldistp = (qlogis(dist_perc+0.001))
# predicting effect of pastoral status:
pastoral.pred.logit = data.frame(pastoral = c(“pastoral”,”pastoral”,”pastoral”,”not.pastoral”,”not.pastoral”,”not.pastoral”),
logit.percent.disturbed = c(-5.48660829,
-5.29961149,
-5.67360509,
-4.94659229,
-4.75959549,
-5.13358909))
pastoral.pred.logit$percent.disturbed = plogis(pastoral.pred.logit$logit.percent.disturbed)-0.001
pastoral.pred.logit$percent.disturbed
pastoral.pred2 fit = (c(0.0031247841,0.0060574270)*100))
pastoral.pred2$se.low = (c((0.003124784-0.002423696),(0.006057427-0.004860812))*100)
pastoral.pred2$se.high = (c((0.003968722-0.003124784),(0.007496270-0.006057427))*100)
k k + geom_pointrange(size=1,aes(colour=Pastoral.status)) + theme_bw()+ylab(“Percent of area disturbed”)
ggsave(“dp figure 1 – pastoral status.png”, width=4, height=3, dpi=300)
# using intervals function to calculate predictions and 95% intervals:
conf.ints = intervals(finaldp)
# predicting effect of pastoral status:
pastoral.pred1 = data.frame(pastoral = c(“pastoral”,”pastoral”,”pastoral”,”not.pastoral”,”not.pastoral”,”not.pastoral”),
logit.percent.disturbed = c(-5.40035707,
-3.81570717,
-6.98500697,
-4.86034100,
-3.30078471,
-6.41989730))
pastoral.pred1$percent.disturbed = plogis(pastoral.pred1$logit.percent.disturbed)-0.001
attach(pastoral.pred1)
fit1=percent.disturbed[1];low1 = percent.disturbed[3];high1=percent.disturbed[2]
fit2=percent.disturbed[4];low2 = percent.disturbed[6];high2=percent.disturbed[5]
pastoral.pred2 fit = c(fit1,fit2))
pastoral.pred2$se.low = c((fit1-low1),(fit2-low2))
pastoral.pred2$se.high = c((high1-fit1),(high2-fit2))
k k + geom_pointrange(size=1,aes(colour=Pastoral.status)) + theme_bw()+ylab(“Percent of area disturbed”)
ggsave(“dp figure 1 – pastoral status(full uncertainty).png”, width=4, height=3, dpi=300)
# predicting effect of esa status:
esa.pred1 = data.frame(esa = c(“esa”,”esa”,”esa”,”not.esa”,”not.esa”,”not.esa”),
logit.percent.disturbed = c(-3.83612729,
-3.64913049,
-4.02312409,
-4.94659229,
-4.75959549,
-5.13358909))
esa.pred1$percent.disturbed = (plogis(esa.pred1$logit.percent.disturbed)-0.001)*100
attach(esa.pred1)
fit1=percent.disturbed[1];low1 = percent.disturbed[3];high1=percent.disturbed[2]
fit2=percent.disturbed[4];low2 = percent.disturbed[6];high2=percent.disturbed[5]
esa.pred2 fit = c(fit1,fit2))
esa.pred2$se.low = c((fit1-low1),(fit2-low2))
esa.pred2$se.high = c((high1-fit1),(high2-fit2))
k k + geom_pointrange(size=1,aes(colour=Clearing.regs)) + theme_bw()+ylab(“Percent of area disturbed”)
ggsave(“dp figure 2 – esa.png”, width=4, height=3, dpi=300)
# predicting effect of conservation estate:
table(dp$cons_estate)
cons.pred1 = data.frame(cons = c(rep(c(“class-A”,”ex.leasehold”,”gazetted”,”not-cons”,”unofficial”),each=3)),
logit.percent.disturbed = c(-5.70267029, -4.91130149, -6.49403909,
-4.34291529, -3.60996819, -5.07586239,
-6.59664929, -5.88199979, -7.31129879,
-4.94659229, -4.30305129, -5.59013329,
-5.45534029, -4.75924019, -6.15144039))
cons.pred1$percent.disturbed = plogis(cons.pred1$logit.percent.disturbed)-0.001
attach(cons.pred1)
fit1= percent.disturbed[1]; low1 = percent.disturbed[3]; high1=percent.disturbed[2]
fit2= percent.disturbed[4]; low2 = percent.disturbed[6]; high2=percent.disturbed[5]
fit3= percent.disturbed[7]; low3 = percent.disturbed[9]; high3=percent.disturbed[8]
fit4= percent.disturbed[10];low4 = percent.disturbed[12];high4=percent.disturbed[11]
fit5= percent.disturbed[13];low5 = percent.disturbed[15];high5=percent.disturbed[14]
cons.pred2 fit = c(fit1,fit2,fit3,fit4,fit5))
cons.pred2$se.low = c((fit1-low1),(fit2-low2),(fit3-low3),(fit4-low4),(fit5-low5))
cons.pred2$se.high = c((high1-fit1),(high2-fit2),(high3-fit3),(high4-fit4),(high5-fit5))
k k + geom_pointrange(size=1,aes(colour=Conservation.tenure)) + theme_bw()+ylab(“Percent of area disturbed”)
ggsave(“dp figure 3 – cons.png”, width=6, height=3, dpi=300) # note colouring is different to combined plot (r2 code) but content is same
# predicting effect of vegetation form:
table(dp$veg_form)
veg.pred1 = data.frame(veg = c(rep(c(“bare”,”broombush”,”mallee”,”mulga”,”shrubland”,”succulent”,”woodland”),each=3)),
logit.percent.disturbed = c(-4.56203229, -5.35340109, -3.77066349,
-4.12225829, -4.35071239, -3.89380419,
-4.33060829, -4.71315739, -3.94805919,
-3.40862529, -3.71443229, -3.10281829,
-4.10232729, -4.37808419, -3.82657039,
-4.00632829, -4.30158499, -3.71107159,
-3.88034929, -4.08146089, -3.67923769))
veg.pred1$percent.disturbed = plogis(veg.pred1$logit.percent.disturbed)-0.001
attach(veg.pred1)
fit1=percent.disturbed[1];low1 = percent.disturbed[2];high1=percent.disturbed[3]
fit2=percent.disturbed[4];low2 = percent.disturbed[5];high2=percent.disturbed[6]
fit3=percent.disturbed[7];low3 = percent.disturbed[8];high3=percent.disturbed[9]
fit4=percent.disturbed[10];low4 = percent.disturbed[11];high4=percent.disturbed[12]
fit5=percent.disturbed[13];low5 = percent.disturbed[14];high5=percent.disturbed[15]
fit6=percent.disturbed[16];low6 = percent.disturbed[17];high6=percent.disturbed[18]
fit7=percent.disturbed[19];low7 = percent.disturbed[20];high7=percent.disturbed[21]
veg.pred2 fit = c(fit1,fit2,fit3,fit4,fit5,fit6,fit7))
veg.pred2$se.low = c((fit1-low1),(fit2-low2),(fit3-low3),(fit4-low4),(fit5-low5),(fit6-low6),(fit7-low7))
veg.pred2$se.high = c((high1-fit1),(high2-fit2),(high3-fit3),(high4-fit4),(high5-fit5),(high6-fit6),(high7-fit7))
k k + geom_pointrange(size=1,aes(colour=Vegetation.form)) + theme_bw()+ylab(“Percent of area disturbed”)+xlab(“Vegetation formation”)
ggsave(“dp figure 4 – veg.png”, width=6, height=3, dpi=300)
summary(dp$dist_perc~dp$veg_form)
summary(veg.pred$percent.disturbed~ veg.pred$veg)
# predicting effect of greenstone lithology:
summary(dp$dist_perc~dp$greenstone)
green.pred1 = data.frame(green = c(rep(c(“greenstone”,”not_greenstone”),each=3)),
logit.percent.disturbed = c(-4.55671829, -5.34808709, -3.76534949,
-4.94659229, -5.09247909, -4.80070549))
green.pred1$percent.disturbed = plogis(green.pred1$logit.percent.disturbed)-0.001
attach(green.pred1)
fit1=percent.disturbed[1];low1 = percent.disturbed[2];high1=percent.disturbed[3]
fit2=percent.disturbed[4];low2 = percent.disturbed[5];high2=percent.disturbed[6]
green.pred2 fit = c(fit1,fit2))
green.pred2$se.low = c((fit1-low1),(fit2-low2))
green.pred2$se.high = c((high1-fit1),(high2-fit2))
k k + geom_pointrange(size=1,aes(colour=Lithology)) + theme_bw()+ylab(“Percent of area disturbed”)+xlab(“Lithology”)
ggsave(“dp figure 5 – greenstone.png”, width=4, height=3, dpi=300)
# predicting effect of tenement years:
range(lntyrs) # 0.000000 to 5.181784
tenyrs.pred = data.frame(lntyrs = seq(from=0, to= 5.181784, by= 0.01))
tenyrs.pred$logit.percent.disturbed = -5.107206754 + 0.440025 *tenyrs.pred$lntyrs
tenyrs.pred$logit.percent.disturbed.upper = -5.107206754 + (0.440025+0.0756683) *tenyrs.pred$lntyrs
tenyrs.pred$logit.percent.disturbed.lower = -5.107206754 + (0.440025-0.0756683) *tenyrs.pred$lntyrs
tenyrs.pred$tenyrs = exp(tenyrs.pred$lntyrs)-1
tenyrs.pred$percent.disturbed = (plogis(tenyrs.pred$logit.percent.disturbed)-0.001)*100
tenyrs.pred$percent.disturbed.upper = (plogis(tenyrs.pred$logit.percent.disturbed.upper)-0.001)*100
tenyrs.pred$percent.disturbed.lower = (plogis(tenyrs.pred$logit.percent.disturbed.lower)-0.001)*100
plot(percent.disturbed~lntyrs, data = tenyrs.pred, xlab = “log of tenement years”, ylab=”Percentage of area disturbed”)
head(tenyrs.pred) # variables of use are: tenyrs,percent.disturbed,percent.disturbed.upper,percent.disturbed.lower
#start of plot
p = ggplot(tenyrs.pred, aes(tenyrs,percent.disturbed))
p + geom_line(colour=I(“red”),size=1.1)+
geom_ribbon(aes(ymin=percent.disturbed.lower, ymax=percent.disturbed.upper),
fill=I(“red”),alpha=0.2)+
theme_bw() +
labs(x=”Total tenement duration (years)”,y=”Area disturbed (%)”)
ggsave(“dp figure 6 – tenement years.png”, width=4, height=3, dpi=300)
# end of plot
# predicting effect of distance to town:
range(tnkm) # 0.74 100.00
tnkm.pred = data.frame(tnkm = seq(from=0.74, to= 100, by= 0.1))
tnkm.pred$logit.percent.disturbed = -2.629818536 + -0.019183 *tnkm.pred$tnkm
tnkm.pred$logit.percent.disturbed.upper = -2.629818536 + (-0.019183+0.0051545) *tnkm.pred$tnkm
tnkm.pred$logit.percent.disturbed.lower = -2.629818536 + (-0.019183-0.0051545) *tnkm.pred$tnkm
tnkm.pred$percent.disturbed = (plogis(tnkm.pred$logit.percent.disturbed)-0.001)*100
tnkm.pred$percent.disturbed.upper = (plogis(tnkm.pred$logit.percent.disturbed.upper)-0.001)*100
tnkm.pred$percent.disturbed.lower = (plogis(tnkm.pred$logit.percent.disturbed.lower)-0.001)*100
plot(percent.disturbed~tnkm, data = tnkm.pred, xlab = “Distance to town (km)”, ylab=”Percentage of area disturbed”)
#start of plot
p = ggplot(tnkm.pred, aes(tnkm,percent.disturbed))
p + geom_line(colour=I(“blue”),size=1.1)+
geom_ribbon(aes(ymin=percent.disturbed.lower, ymax=percent.disturbed.upper),
fill=I(“blue”),alpha=0.2)+
theme_bw() +
labs(x=”Distance from town (kms)”,y=”Area disturbed (%)”)
ggsave(“dp figure 7 – distance from town.png”, width=4, height=3, dpi=300)
# end of plot
###########################################################################################################
# an afterthought: it would be good to combine pastoral and cons_estate as they both refer to tenure.
# I’ve resolved a few conflicts and created a new variable called ‘tenure’; the AIC for this model is slightly better,
dptenure = lme(ldistp ~ factor(tenure) + factor(esa) + factor(veg_form) + factor(greenstone) +
lntyrs + tnkm, random = ~1|SA_number,method = “REML”, data=dp)
summary(dptenure)
summary(dp$tenure)
# Maybe this would be better but I’d have to do all the predictions over again in excel and that’s too hard
# but for now I’ll have a sneak peak at the result of it:
# Class-A is the default,
# former leasehold has more disturbance
# gazetted cons has less disturbance
# not-cons has more disturbance but less than former leasehold
# pastoral has more disturbance but less than not-cons
# unofficial cons has more than pastoral
# so in order of most to least disturbance:
# 1. former leasehold
# 2. not-cons-estate
# 3. unofficial cons
# 4. pastoral
# 5. Class-A
# 6. gazetted conservation
# It’s wierd, you’d expect pastoral and former-pastoral to match up a bit more.
# ..and perhaps unofficial cons to match up more with gazetted cons.
# it looks as if the process of converting a pastoral lease to a conservation-ex-lease is the most disturbing thing!
# doesn’t really make sense, unless it’s all focued on Credo and they cleared a lot more area?
# I think i need to actually just combine the pastoral and ex-pastoral categories and that might increase the sample size and solve the issue.
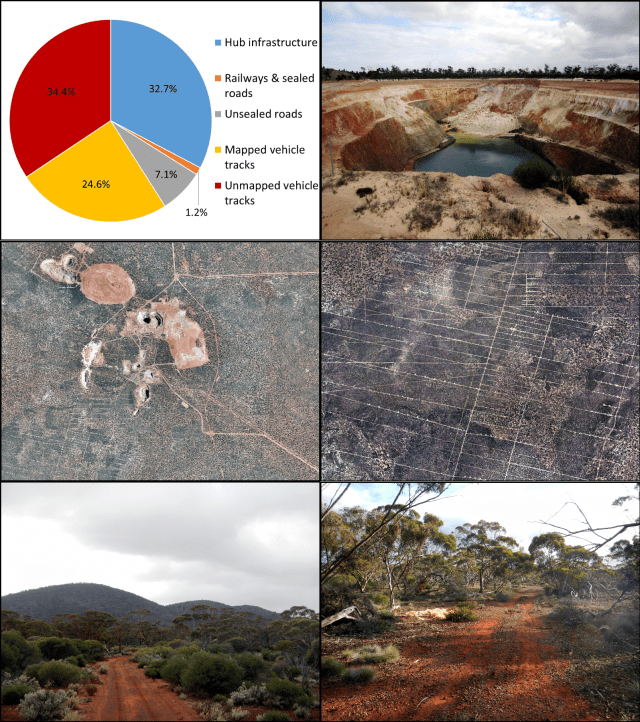

 The Great Western Woodlands
The Great Western Woodlands
 Don’t miss being a part of the inaugural Jungkajungka Woodlands Festival, held over Easter in Norseman, Western Australia—the Heart of the Great Western Woodlands. This event is organised by the Wilderness Society in collaboration with the Shire of Dundas, GondwanaLink, and with support from a number of other organisations.
Don’t miss being a part of the inaugural Jungkajungka Woodlands Festival, held over Easter in Norseman, Western Australia—the Heart of the Great Western Woodlands. This event is organised by the Wilderness Society in collaboration with the Shire of Dundas, GondwanaLink, and with support from a number of other organisations.